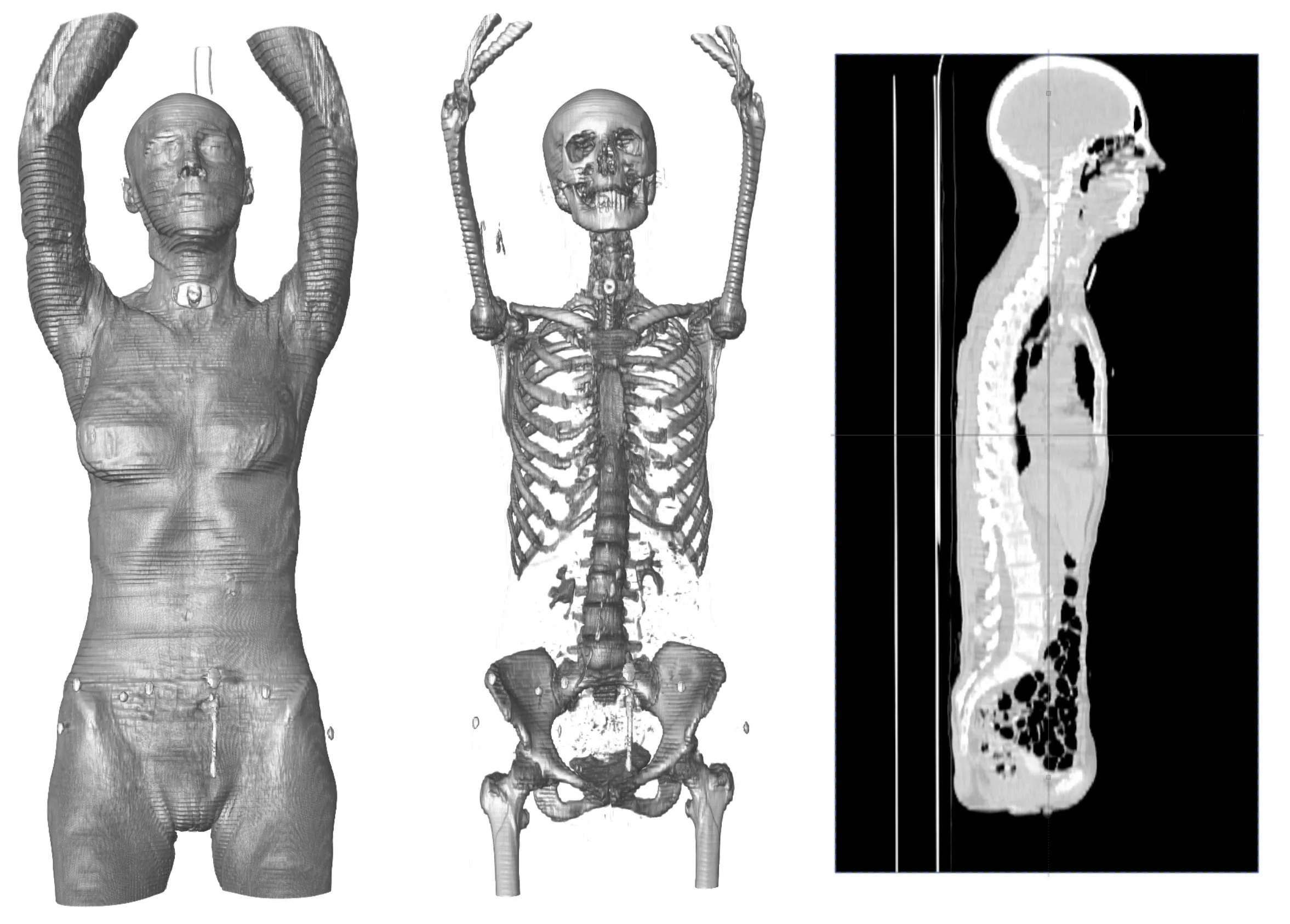
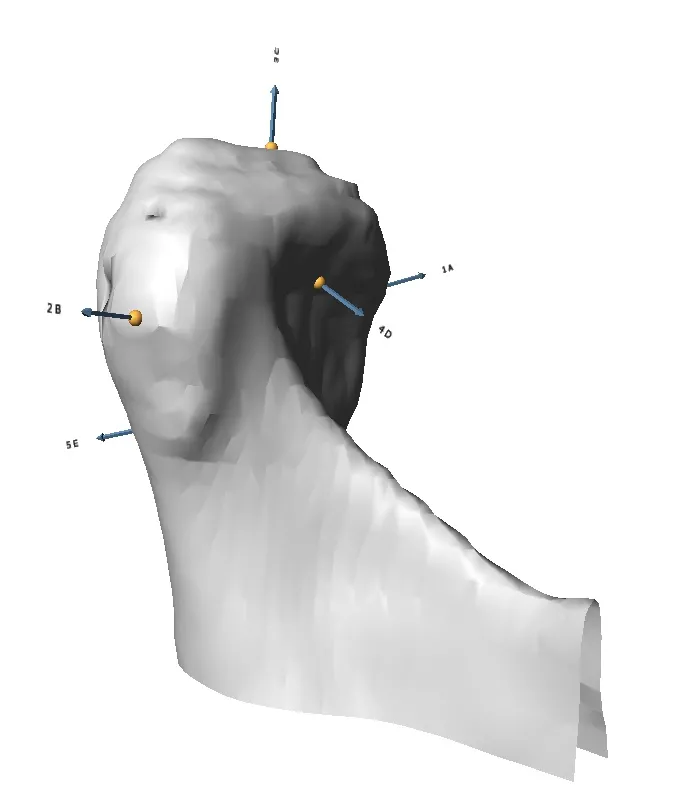
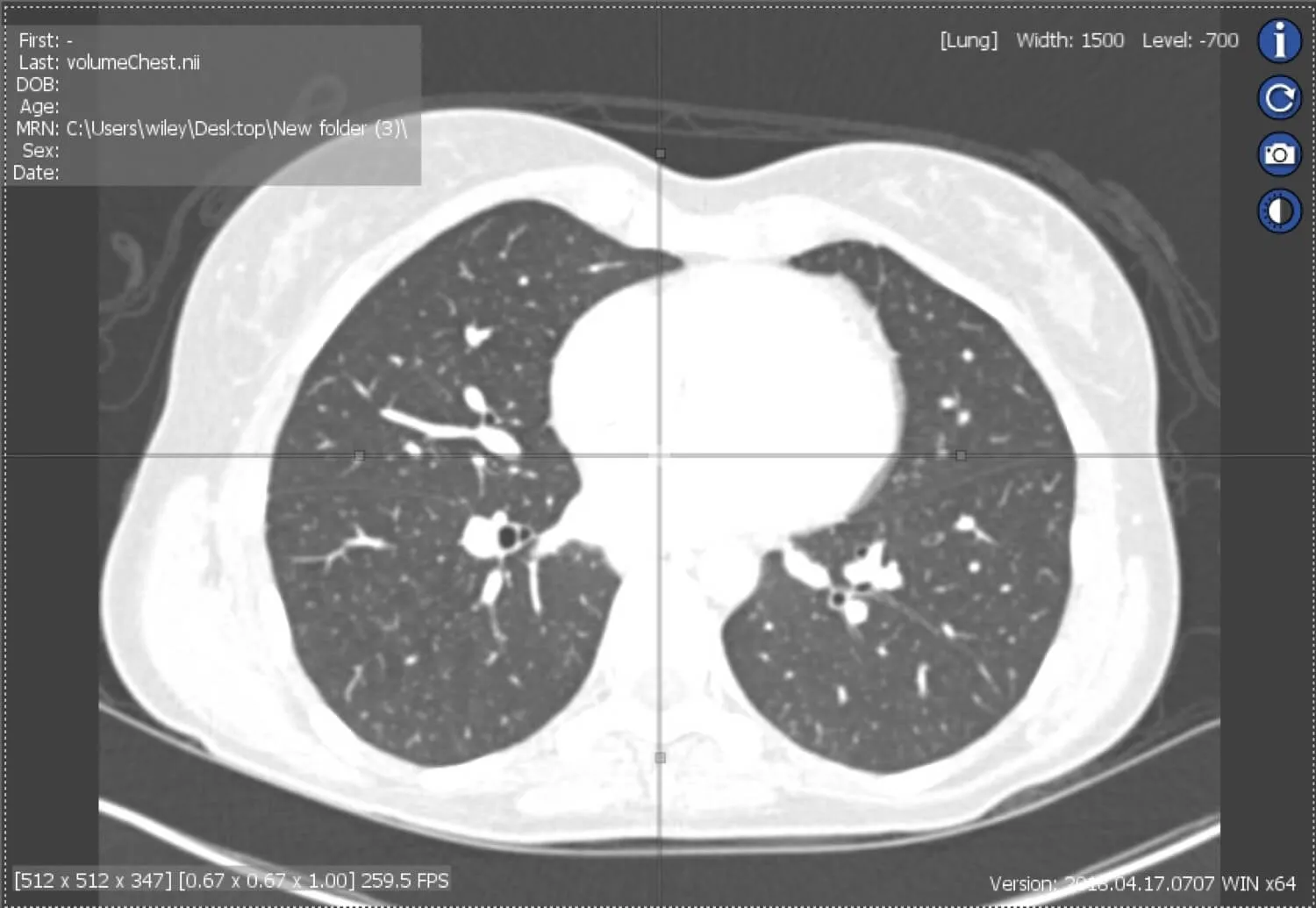
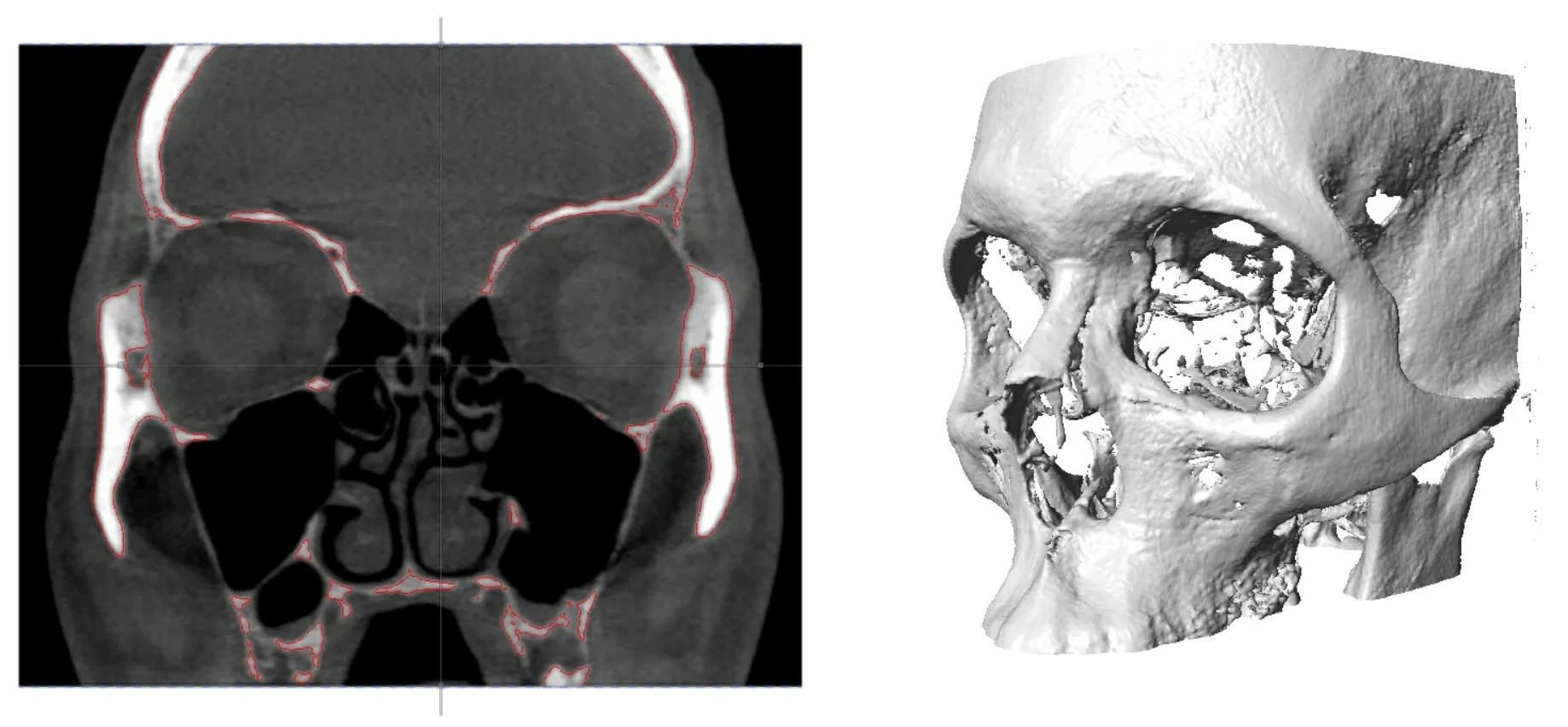
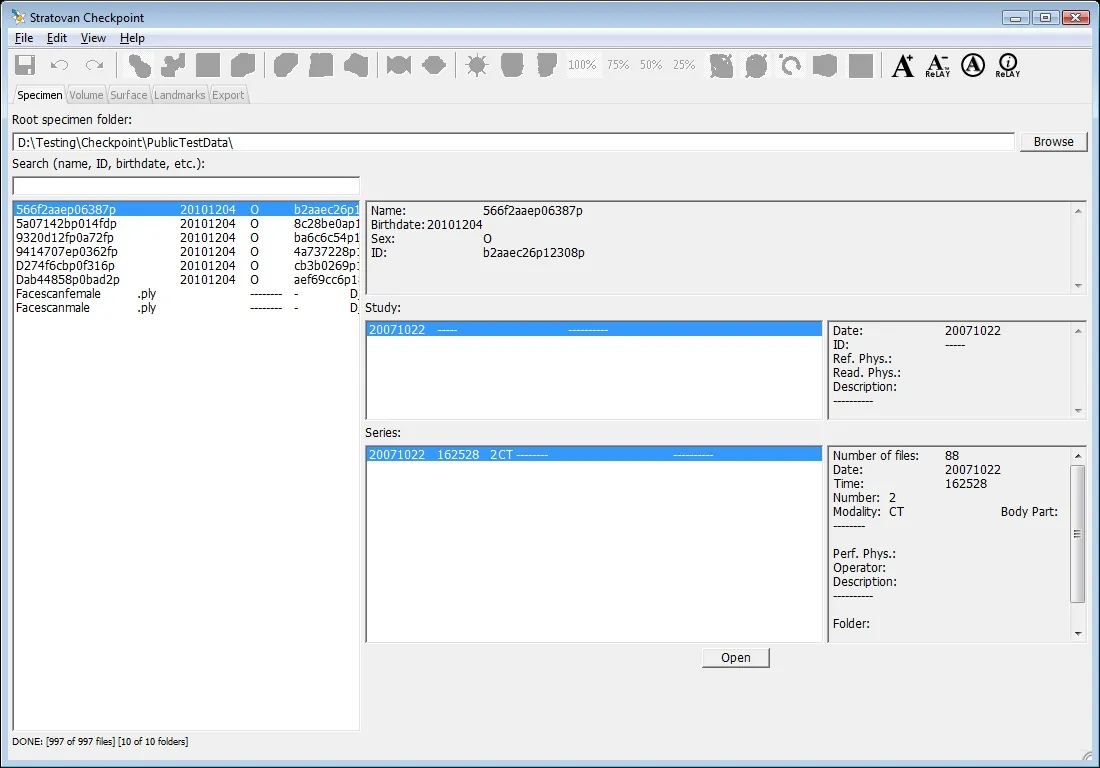
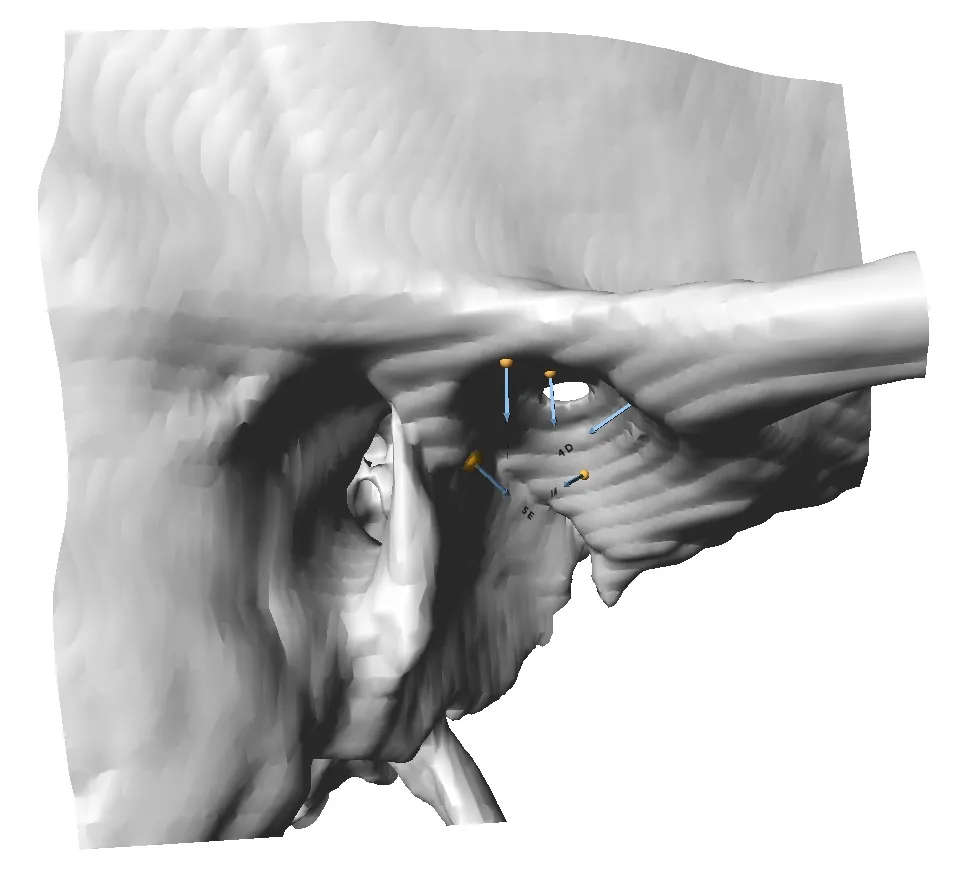
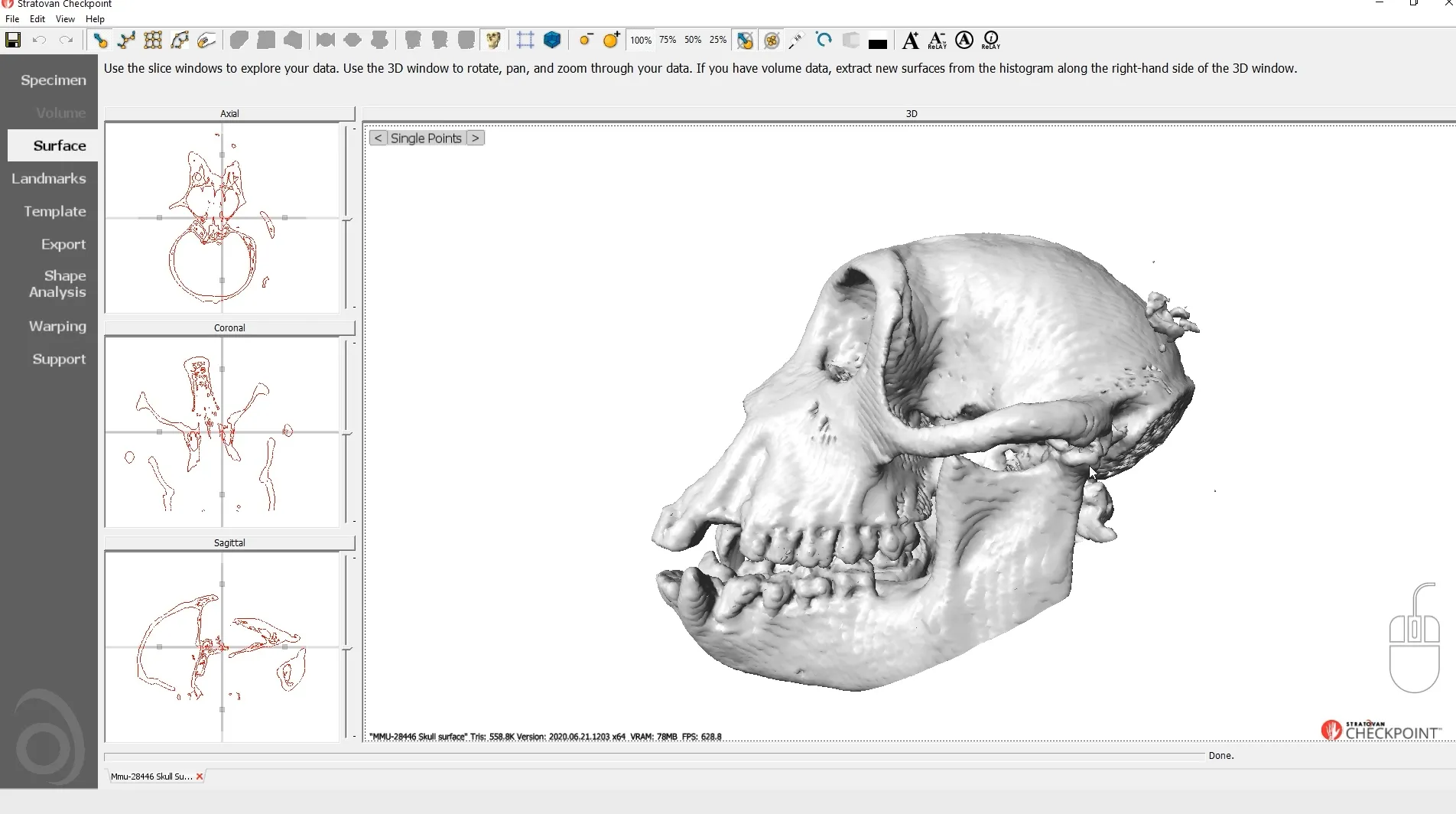
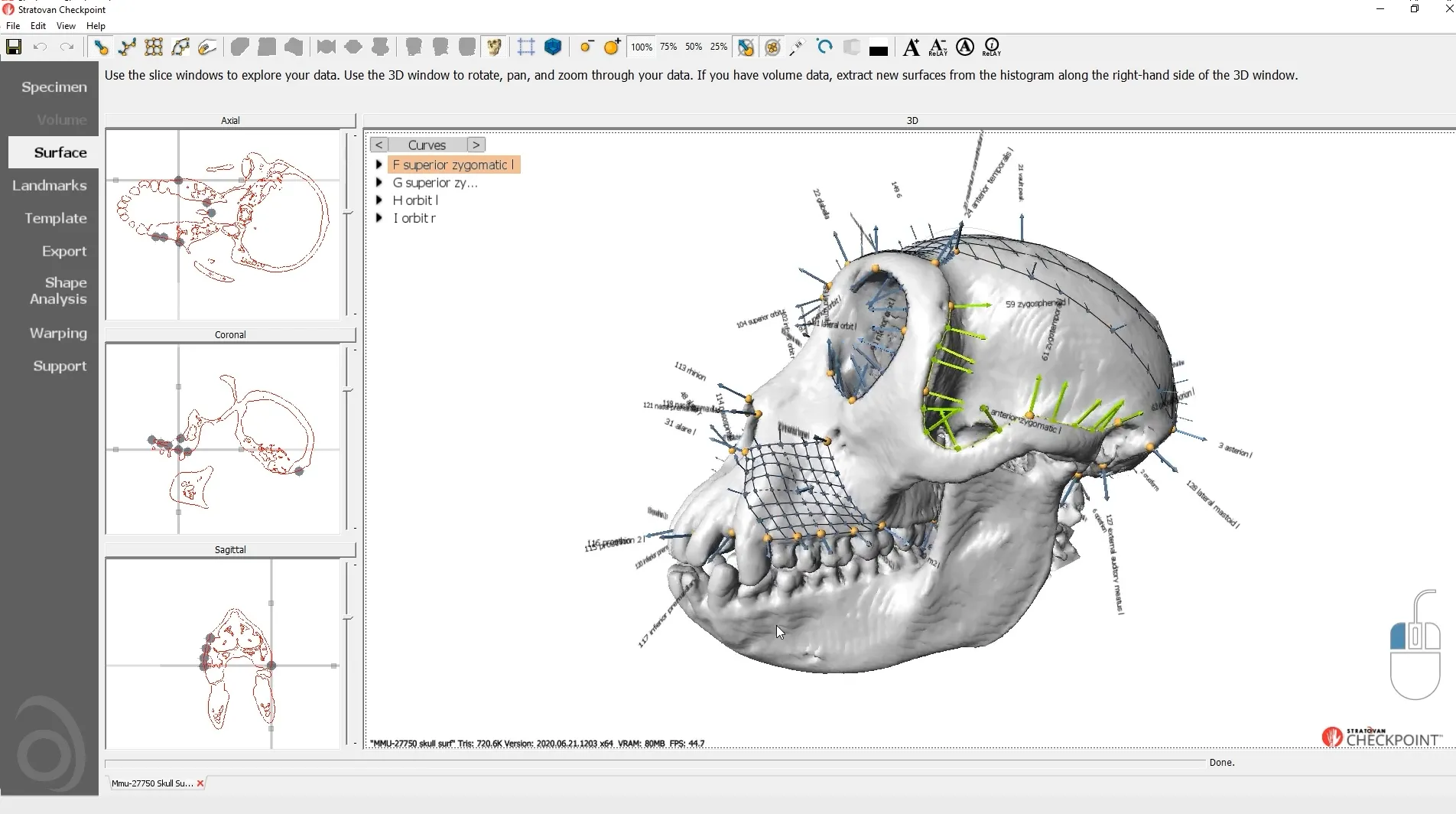
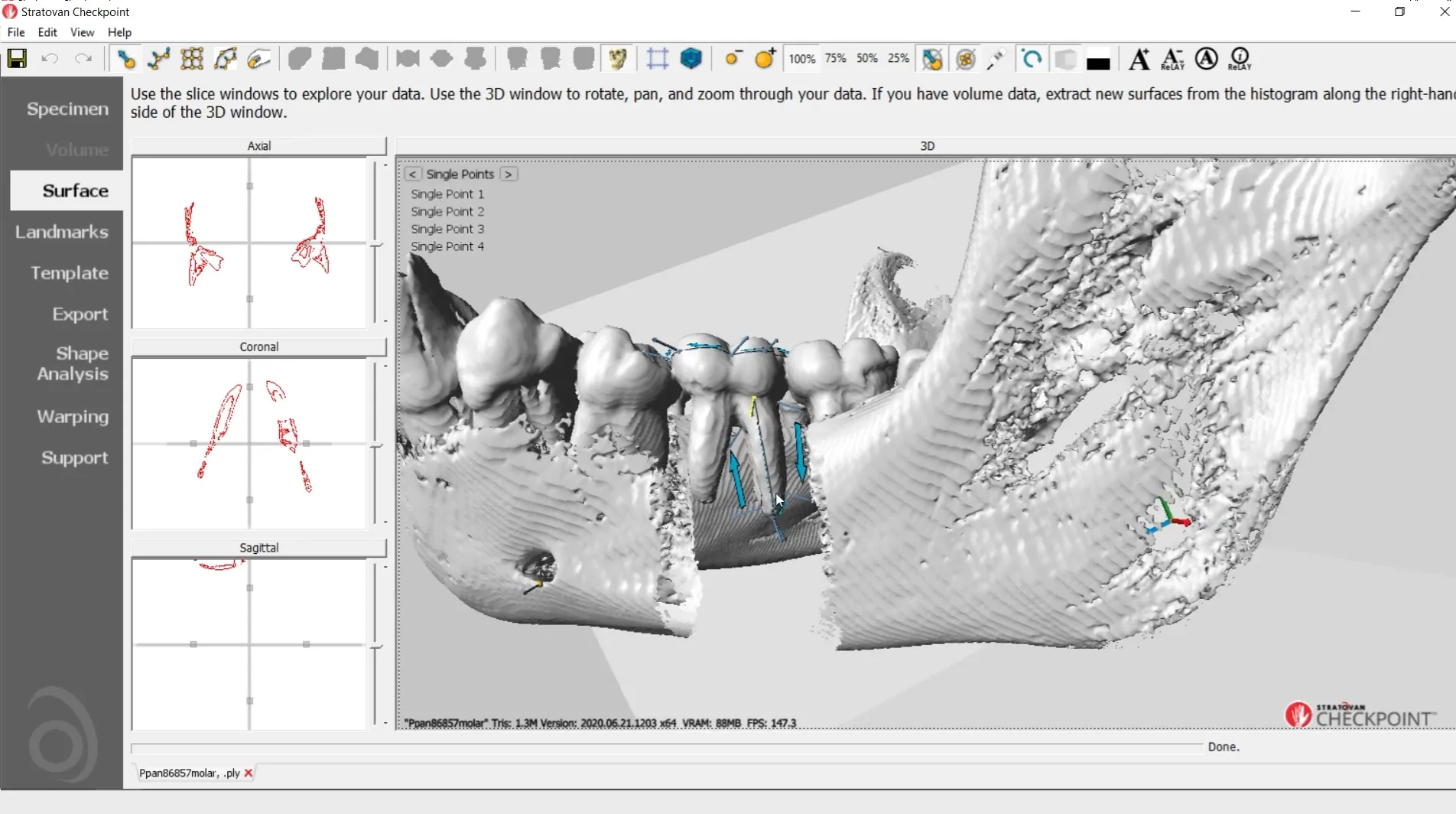
Checkpoint - the most used 3D shape analysis & morphometrics software in the World
Used by thousands of researchers at hundreds of top research institutions and museums around the globe, Checkpoint is an integrated software package for geometric morphometrics that enables you to easily collect, manipulate and analyze thousands of landmark and semi-landmark points in 3D.